As of writing, the answer is...fairly easy, thanks to Wilson Freitas' new rmd_reader plugin.
However, there's a little bit of trickiness regarding figure paths. knitr has amazing functionality for report generation, including the ability to easily embed plots within an .Rmd document.
In reality, though, knitr is generating the plot for you behind the scenes, and when you render your output (e.g. "knit html"), knitr is creating the image and saving it to a directory that is in the output path.
For example, say you have a file "temp.Rmd" that has an embedded plot and you knit the file to HTML. If you don't specify a fig.path for knitr, this will produce a bunch of files (your .md file, your .html file), as well as a "figure" directory. In that directory will be your embedded plot as an image, and your .html file will refer to that path.
So, why does this matter? The rmd_reader pelican plug-in currently doesn't specify how to handle paths. And to be honest, I wouldn't hold my breath for a patch; people can organize their content directory for pelican in any way they want.
For example, my pelican content directory is structured as so:
- content
-- articles
--- tutorials
--- updates
--- walkthroughs
-- notebooks
-- pages
-- images
-- figure
-- extra
In each of those article subdirectories, I want to be able to put an .Rmd file in there that will automatically generate the appropriate article in the right location.
But take a note of my ARTICLE_URL and ARTICLE_SAVE_AS options in pelicanconf.py:
ARTICLE_URL = "articles/{date:%Y}_{date:%m}_{date:%d}/{slug}/"
ARTICLE_SAVE_AS = "articles/{date:%Y}_{date:%m}_{date:%d}/{slug}/index.html"
This means that when I generate a new plot, they're going to be in an articles/date/{slug}/ relative path on my website. And since the "figure" directory is actually in the base directory of the website, we've got a broken link problem.
Here's my dirty fix (which works). Just set your knitr fig.path in your .Rmd appropriately, like so:
opts_chunk$set(fig.path='../../../figure/',echo=FALSE, warning=FALSE, message=FALSE)
Don't forget the trailing / in the fig.path!
And with that global option set, you should be able to continue using knitr as you are used to. Here's the template R Markdown document from RStudio:
Title
This is an R Markdown document. Markdown is a simple formatting syntax for authoring web pages (click the Help toolbar button for more details on using R Markdown).
When you click the Knit HTML button a web page will be generated that includes both content as well as the output of any embedded R code chunks within the document. You can embed an R code chunk like this:
## speed dist
## Min. : 4.0 Min. : 2.00
## 1st Qu.:12.0 1st Qu.: 26.00
## Median :15.0 Median : 36.00
## Mean :15.4 Mean : 42.98
## 3rd Qu.:19.0 3rd Qu.: 56.00
## Max. :25.0 Max. :120.00
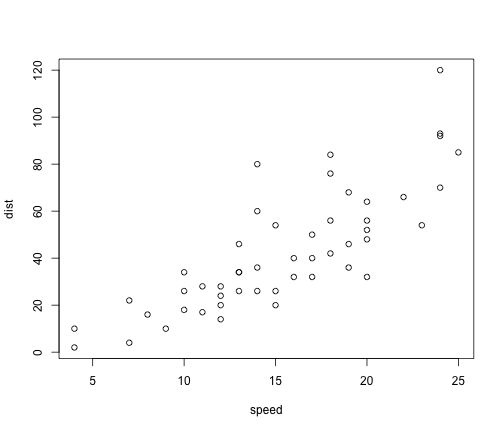
You can also embed plots, for example:
 Rebecca Weiss
Rebecca Weiss
Comments
comments powered by Disqus